step_impute_linear() creates a specification of a recipe step that will
create linear regression models to impute missing data.
Usage
step_impute_linear(
recipe,
...,
role = NA,
trained = FALSE,
impute_with = all_predictors(),
models = NULL,
skip = FALSE,
id = rand_id("impute_linear")
)Arguments
- recipe
A recipe object. The step will be added to the sequence of operations for this recipe.
- ...
One or more selector functions to choose variables to be imputed; these variables must be of type
numeric. When used withimp_vars, these dots indicate which variables are used to predict the missing data in each variable. Seeselections()for more details.- role
Not used by this step since no new variables are created.
- trained
A logical to indicate if the quantities for preprocessing have been estimated.
- impute_with
Bare names or selectors functions that specify which variables are used to impute the variables that can include specific variable names separated by commas or different selectors (see
selections()). If a column is included in both lists to be imputed and to be an imputation predictor, it will be removed from the latter and not used to impute itself.- models
The
lm()objects are stored here once the linear models have been trained byprep().- skip
A logical. Should the step be skipped when the recipe is baked by
bake()? While all operations are baked whenprep()is run, some operations may not be able to be conducted on new data (e.g. processing the outcome variable(s)). Care should be taken when usingskip = TRUEas it may affect the computations for subsequent operations.- id
A character string that is unique to this step to identify it.
Value
An updated version of recipe with the new step added to the
sequence of any existing operations.
Details
For each variable requiring imputation, a linear model is fit where the
outcome is the variable of interest and the predictors are any other
variables listed in the impute_with formula. Note that if a variable that
is to be imputed is also in impute_with, this variable will be ignored.
The variable(s) to be imputed must be of type numeric. The imputed values
will keep the same type as their original data (i.e, model predictions are
coerced to integer as needed).
Since this is a linear regression, the imputation model only uses complete cases for the training set predictors.
Tidying
When you tidy() this step, a tibble is returned with
columns terms, model , and id:
- terms
character, the selectors or variables selected
- model
list, list of fitted
lm()models- id
character, id of this step
Case weights
This step performs an unsupervised operation that can utilize case weights.
As a result, case weights are only used with frequency weights. For more
information, see the documentation in case_weights and the examples on
tidymodels.org.
References
Kuhn, M. and Johnson, K. (2013). Feature Engineering and Selection https://feat.engineering/08-Handling_Missing_Data.html
See also
Other imputation steps:
step_impute_bag(),
step_impute_knn(),
step_impute_lower(),
step_impute_mean(),
step_impute_median(),
step_impute_mode(),
step_impute_roll()
Examples
data(ames, package = "modeldata")
set.seed(393)
ames_missing <- ames
ames_missing$Longitude[sample(1:nrow(ames), 200)] <- NA
imputed_ames <-
recipe(Sale_Price ~ ., data = ames_missing) |>
step_impute_linear(
Longitude,
impute_with = c(Latitude, Neighborhood, MS_Zoning, Alley)
) |>
prep(ames_missing)
imputed <-
bake(imputed_ames, new_data = ames_missing) |>
dplyr::rename(imputed = Longitude) |>
bind_cols(ames |> dplyr::select(original = Longitude)) |>
bind_cols(ames_missing |> dplyr::select(Longitude)) |>
dplyr::filter(is.na(Longitude))
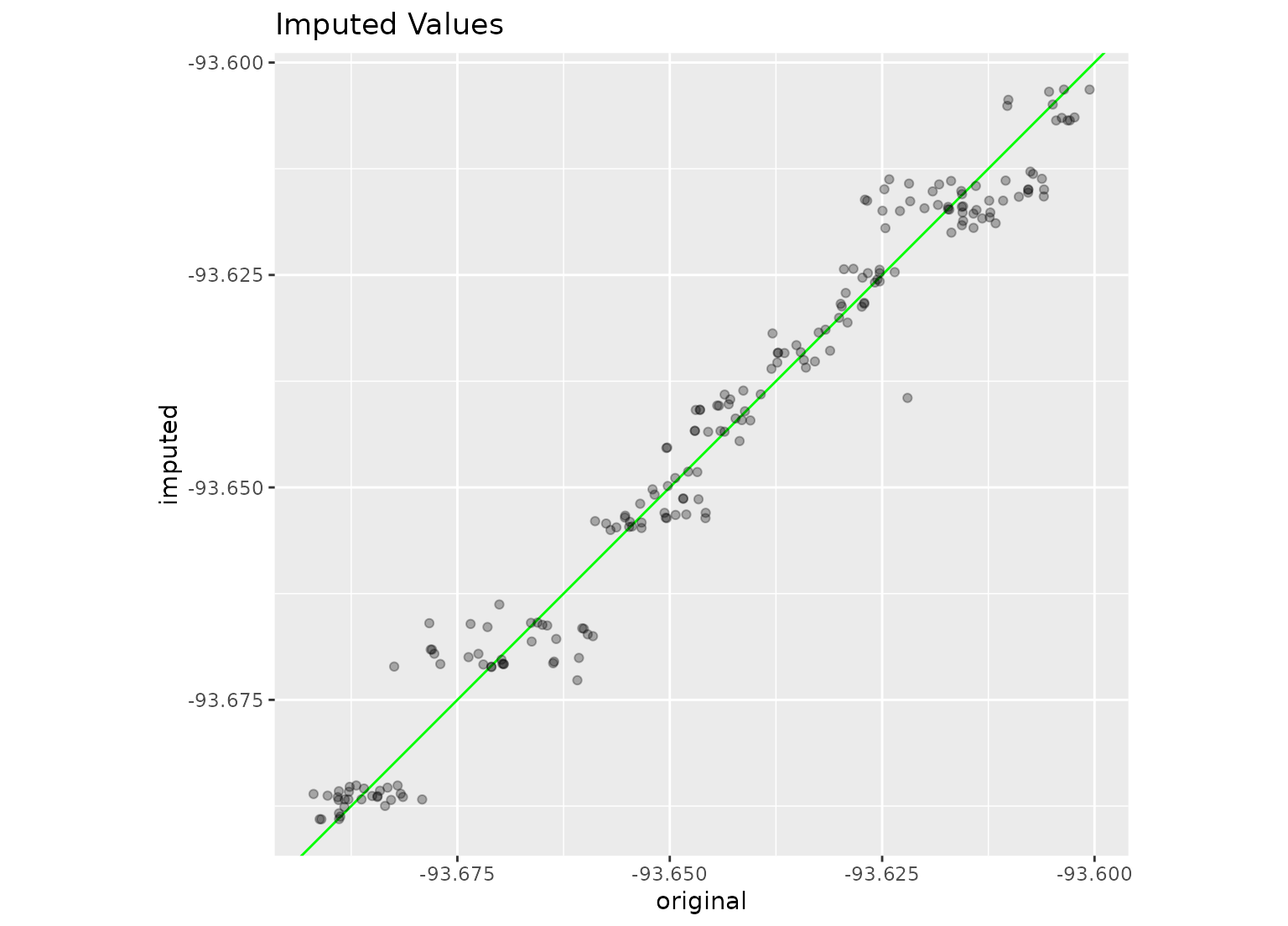
library(ggplot2)
ggplot(imputed, aes(x = original, y = imputed)) +
geom_abline(col = "green") +
geom_point(alpha = .3) +
coord_equal() +
labs(title = "Imputed Values")