step_kpca() creates a specification of a recipe step that will convert
numeric data into one or more principal components using a kernel basis
expansion.
Arguments
- recipe
A recipe object. The step will be added to the sequence of operations for this recipe.
- ...
One or more selector functions to choose variables for this step. See
selections()for more details.- role
For model terms created by this step, what analysis role should they be assigned? By default, the new columns created by this step from the original variables will be used as predictors in a model.
- trained
A logical to indicate if the quantities for preprocessing have been estimated.
- num_comp
The number of components to retain as new predictors. If
num_compis greater than the number of columns or the number of possible components, a smaller value will be used. Ifnum_comp = 0is set then no transformation is done and selected variables will stay unchanged, regardless of the value ofkeep_original_cols.- res
An S4
kernlab::kpca()object is stored here once this preprocessing step has be trained byprep().- columns
A character string of the selected variable names. This field is a placeholder and will be populated once
prep()is used.- options
A list of options to
kernlab::kpca(). Defaults are set for the argumentskernelandkparbut others can be passed in. Note that the argumentsxandfeaturesshould not be passed here (or at all).- prefix
A character string for the prefix of the resulting new variables. See notes below.
- keep_original_cols
A logical to keep the original variables in the output. Defaults to
FALSE.- skip
A logical. Should the step be skipped when the recipe is baked by
bake()? While all operations are baked whenprep()is run, some operations may not be able to be conducted on new data (e.g. processing the outcome variable(s)). Care should be taken when usingskip = TRUEas it may affect the computations for subsequent operations.- id
A character string that is unique to this step to identify it.
Value
An updated version of recipe with the new step added to the
sequence of any existing operations.
Details
When performing kPCA with step_kpca(), you must choose the kernel function
(and any important kernel parameters). This step uses the kernlab
package; the reference below discusses the types of kernels available and
their parameter(s). These specifications can be made in the kernel and
kpar slots of the options argument to step_kpca(). Consider using
step_kpca_rbf() for a radial basis function kernel or step_kpca_poly()
for a polynomial kernel.
Kernel principal component analysis (kPCA) is an extension of a PCA analysis that conducts the calculations in a broader dimensionality defined by a kernel function. For example, if a quadratic kernel function were used, each variable would be represented by its original values as well as its square. This nonlinear mapping is used during the PCA analysis and can potentially help find better representations of the original data.
This step requires the kernlab package. If not installed, the step will stop with a prompt about installing the package.
As with ordinary PCA, it is important to center and scale the variables
prior to computing PCA components (step_normalize() can be used for
this purpose).
The argument num_comp controls the number of components that will be retained
(the original variables that are used to derive the components are removed from
the data). The new components will have names that begin with prefix and a
sequence of numbers. The variable names are padded with zeros. For example, if
num_comp < 10, their names will be kPC1 - kPC9. If num_comp = 101,
the names would be kPC001 - kPC101.
tidy() results
When you tidy() this step, a tibble with column
terms (the selectors or variables selected) is returned.
Tidying
When you tidy() this step, a tibble is returned with
columns terms and id:
- terms
character, the selectors or variables selected
- id
character, id of this step
References
Scholkopf, B., Smola, A., and Muller, K. (1997). Kernel principal component analysis. Lecture Notes in Computer Science, 1327, 583-588.
Karatzoglou, K., Smola, A., Hornik, K., and Zeileis, A. (2004). kernlab - An S4 package for kernel methods in R. Journal of Statistical Software, 11(1), 1-20.
See also
Other multivariate transformation steps:
step_classdist(),
step_classdist_shrunken(),
step_depth(),
step_geodist(),
step_ica(),
step_isomap(),
step_kpca_poly(),
step_kpca_rbf(),
step_mutate_at(),
step_nnmf(),
step_nnmf_sparse(),
step_pca(),
step_pls(),
step_ratio(),
step_spatialsign()
Examples
library(ggplot2)
data(biomass, package = "modeldata")
biomass_tr <- biomass[biomass$dataset == "Training", ]
biomass_te <- biomass[biomass$dataset == "Testing", ]
rec <- recipe(
HHV ~ carbon + hydrogen + oxygen + nitrogen + sulfur,
data = biomass_tr
)
kpca_trans <- rec |>
step_YeoJohnson(all_numeric_predictors()) |>
step_normalize(all_numeric_predictors()) |>
step_kpca(all_numeric_predictors())
kpca_estimates <- prep(kpca_trans, training = biomass_tr)
kpca_te <- bake(kpca_estimates, biomass_te)
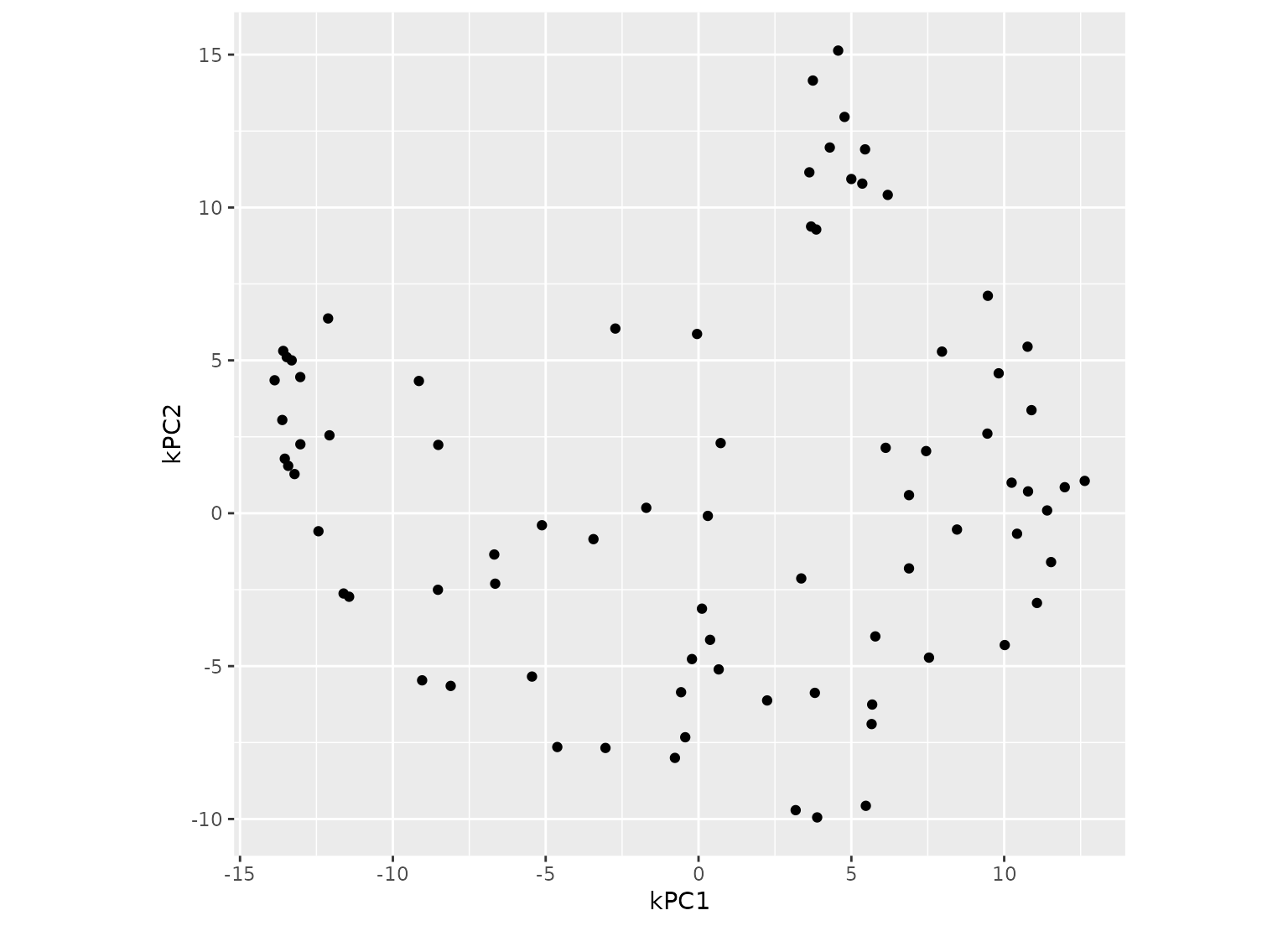
ggplot(kpca_te, aes(x = kPC1, y = kPC2)) +
geom_point() +
coord_equal()
 tidy(kpca_trans, number = 3)
#> # A tibble: 1 × 2
#> terms id
#> <chr> <chr>
#> 1 all_numeric_predictors() kpca_ntsLB
tidy(kpca_estimates, number = 3)
#> # A tibble: 5 × 2
#> terms id
#> <chr> <chr>
#> 1 carbon kpca_ntsLB
#> 2 hydrogen kpca_ntsLB
#> 3 oxygen kpca_ntsLB
#> 4 nitrogen kpca_ntsLB
#> 5 sulfur kpca_ntsLB
tidy(kpca_trans, number = 3)
#> # A tibble: 1 × 2
#> terms id
#> <chr> <chr>
#> 1 all_numeric_predictors() kpca_ntsLB
tidy(kpca_estimates, number = 3)
#> # A tibble: 5 × 2
#> terms id
#> <chr> <chr>
#> 1 carbon kpca_ntsLB
#> 2 hydrogen kpca_ntsLB
#> 3 oxygen kpca_ntsLB
#> 4 nitrogen kpca_ntsLB
#> 5 sulfur kpca_ntsLB
