step_YeoJohnson() creates a specification of a recipe step that will
transform data using a Yeo-Johnson transformation.
Arguments
- recipe
A recipe object. The step will be added to the sequence of operations for this recipe.
- ...
One or more selector functions to choose variables for this step. See
selections()for more details.- role
Not used by this step since no new variables are created.
- trained
A logical to indicate if the quantities for preprocessing have been estimated.
- lambdas
A numeric vector of transformation values. This is
NULLuntil computed byprep().- limits
A length 2 numeric vector defining the range to compute the transformation parameter lambda.
- num_unique
An integer where data that have less possible values will not be evaluated for a transformation.
- na_rm
A logical value indicating whether
NAvalues should be removed during computations.- skip
A logical. Should the step be skipped when the recipe is baked by
bake()? While all operations are baked whenprep()is run, some operations may not be able to be conducted on new data (e.g. processing the outcome variable(s)). Care should be taken when usingskip = TRUEas it may affect the computations for subsequent operations.- id
A character string that is unique to this step to identify it.
Value
An updated version of recipe with the new step added to the
sequence of any existing operations.
Details
The Yeo-Johnson transformation is very similar to the Box-Cox but does not require the input variables to be strictly positive. In the package, the partial log-likelihood function is directly optimized within a reasonable set of transformation values (which can be changed by the user).
This transformation is typically done on the outcome variable using the residuals for a statistical model (such as ordinary least squares). Here, a simple null model (intercept only) is used to apply the transformation to the predictor variables individually. This can have the effect of making the variable distributions more symmetric.
If the transformation parameters are estimated to be very closed to the
bounds, or if the optimization fails, a value of NA is used and no
transformation is applied.
Tidying
When you tidy() this step, a tibble is returned with
columns terms, value , and id:
- terms
character, the selectors or variables selected
- value
numeric, the lambda estimate
- id
character, id of this step
References
Yeo, I. K., and Johnson, R. A. (2000). A new family of power transformations to improve normality or symmetry. Biometrika.
See also
Other individual transformation steps:
step_BoxCox(),
step_bs(),
step_harmonic(),
step_hyperbolic(),
step_inverse(),
step_invlogit(),
step_log(),
step_logit(),
step_mutate(),
step_ns(),
step_percentile(),
step_poly(),
step_relu(),
step_sqrt()
Examples
data(biomass, package = "modeldata")
biomass_tr <- biomass[biomass$dataset == "Training", ]
biomass_te <- biomass[biomass$dataset == "Testing", ]
rec <- recipe(
HHV ~ carbon + hydrogen + oxygen + nitrogen + sulfur,
data = biomass_tr
)
yj_transform <- step_YeoJohnson(rec, all_numeric())
yj_estimates <- prep(yj_transform, training = biomass_tr)
yj_te <- bake(yj_estimates, biomass_te)
plot(density(biomass_te$sulfur), main = "before")
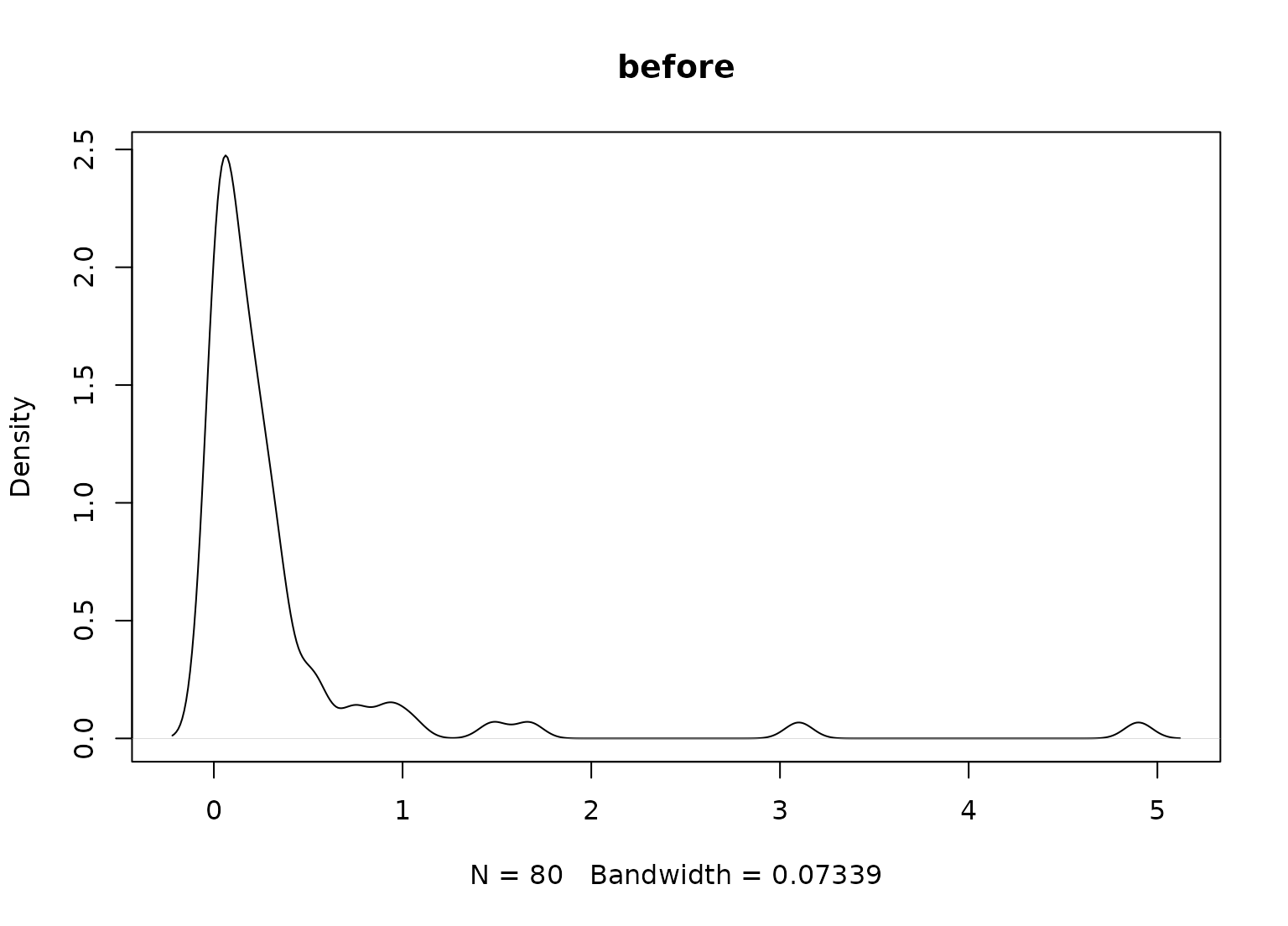 plot(density(yj_te$sulfur), main = "after")
plot(density(yj_te$sulfur), main = "after")
 tidy(yj_transform, number = 1)
#> # A tibble: 1 × 3
#> terms value id
#> <chr> <dbl> <chr>
#> 1 all_numeric() NA YeoJohnson_qjRmV
tidy(yj_estimates, number = 1)
#> # A tibble: 6 × 3
#> terms value id
#> <chr> <dbl> <chr>
#> 1 carbon -0.0225 YeoJohnson_qjRmV
#> 2 hydrogen 2.10 YeoJohnson_qjRmV
#> 3 oxygen 1.78 YeoJohnson_qjRmV
#> 4 nitrogen -0.830 YeoJohnson_qjRmV
#> 5 sulfur -4.09 YeoJohnson_qjRmV
#> 6 HHV -0.388 YeoJohnson_qjRmV
tidy(yj_transform, number = 1)
#> # A tibble: 1 × 3
#> terms value id
#> <chr> <dbl> <chr>
#> 1 all_numeric() NA YeoJohnson_qjRmV
tidy(yj_estimates, number = 1)
#> # A tibble: 6 × 3
#> terms value id
#> <chr> <dbl> <chr>
#> 1 carbon -0.0225 YeoJohnson_qjRmV
#> 2 hydrogen 2.10 YeoJohnson_qjRmV
#> 3 oxygen 1.78 YeoJohnson_qjRmV
#> 4 nitrogen -0.830 YeoJohnson_qjRmV
#> 5 sulfur -4.09 YeoJohnson_qjRmV
#> 6 HHV -0.388 YeoJohnson_qjRmV
