step_hyperbolic() creates a specification of a recipe step that will
transform data using a hyperbolic function.
Arguments
- recipe
A recipe object. The step will be added to the sequence of operations for this recipe.
- ...
One or more selector functions to choose variables for this step. See
selections()for more details.- role
Not used by this step since no new variables are created.
- trained
A logical to indicate if the quantities for preprocessing have been estimated.
- func
A character value for the function. Valid values are
"sinh","cosh", or"tanh".- inverse
A logical: should the inverse function be used?
- columns
A character string of the selected variable names. This field is a placeholder and will be populated once
prep()is used.- skip
A logical. Should the step be skipped when the recipe is baked by
bake()? While all operations are baked whenprep()is run, some operations may not be able to be conducted on new data (e.g. processing the outcome variable(s)). Care should be taken when usingskip = TRUEas it may affect the computations for subsequent operations.- id
A character string that is unique to this step to identify it.
Value
An updated version of recipe with the new step added to the
sequence of any existing operations.
Tidying
When you tidy() this step, a tibble is returned with
columns terms, inverse, func , and id:
- terms
character, the selectors or variables selected
- inverse
logical, is the inverse function be used
- func
character, name of function.
"sinh","cosh", or"tanh"- id
character, id of this step
See also
Other individual transformation steps:
step_BoxCox(),
step_YeoJohnson(),
step_bs(),
step_harmonic(),
step_inverse(),
step_invlogit(),
step_log(),
step_logit(),
step_mutate(),
step_ns(),
step_percentile(),
step_poly(),
step_relu(),
step_sqrt()
Examples
set.seed(313)
examples <- matrix(rnorm(40), ncol = 2)
examples <- as.data.frame(examples)
rec <- recipe(~ V1 + V2, data = examples)
cos_trans <- rec |>
step_hyperbolic(
all_numeric_predictors(),
func = "cosh", inverse = FALSE
)
cos_obj <- prep(cos_trans, training = examples)
transformed_te <- bake(cos_obj, examples)
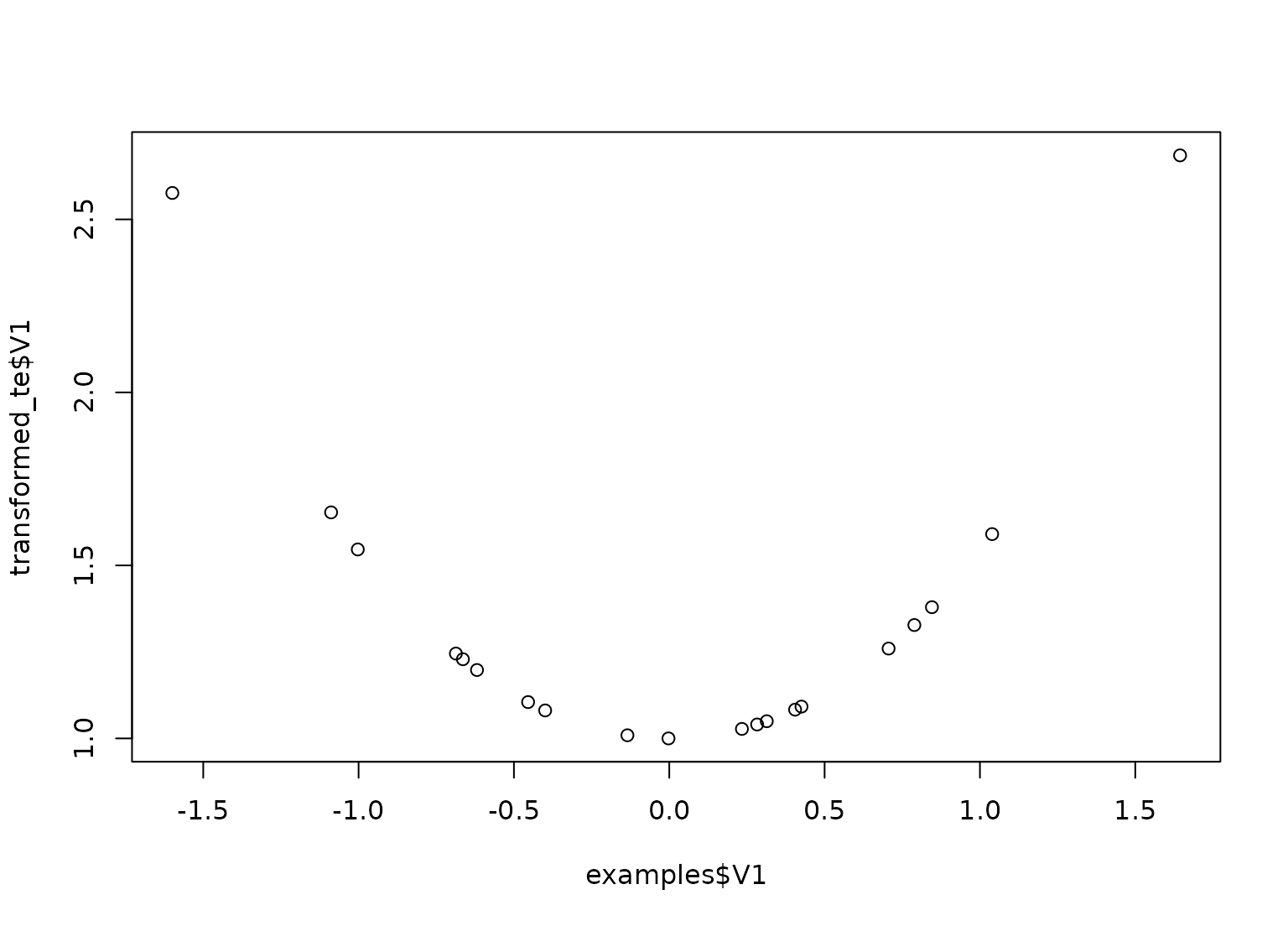
plot(examples$V1, transformed_te$V1)
 tidy(cos_trans, number = 1)
#> # A tibble: 1 × 4
#> terms inverse func id
#> <chr> <lgl> <chr> <chr>
#> 1 all_numeric_predictors() FALSE cosh hyperbolic_IhS7o
tidy(cos_obj, number = 1)
#> # A tibble: 2 × 4
#> terms inverse func id
#> <chr> <lgl> <chr> <chr>
#> 1 V1 FALSE cosh hyperbolic_IhS7o
#> 2 V2 FALSE cosh hyperbolic_IhS7o
tidy(cos_trans, number = 1)
#> # A tibble: 1 × 4
#> terms inverse func id
#> <chr> <lgl> <chr> <chr>
#> 1 all_numeric_predictors() FALSE cosh hyperbolic_IhS7o
tidy(cos_obj, number = 1)
#> # A tibble: 2 × 4
#> terms inverse func id
#> <chr> <lgl> <chr> <chr>
#> 1 V1 FALSE cosh hyperbolic_IhS7o
#> 2 V2 FALSE cosh hyperbolic_IhS7o
